Paul Higgs - Research Topics
RNA Structure and Evolution
Publications
1. N. J. Savill, D.C. Hoyle and P.G. Higgs, P.G. RNA sequence evolution with secondary structure constraints: Comparison of substitution rate models using maximum likelihood methods. Genetics 157, 399-411 (2001).
2. Higgs, P.G. (2000) RNA Secondary Structure: Physical and Computational Aspects. Quart. Rev. Biophys. 33, 199-253.
3. Morgan, S.R. & Higgs, P.G. (1998). Barrier heights between groundstates in a model of RNA secondary structure. J. Phys A (Math. & Gen.) 31, 3153-3170.
4. Higgs, P.G. (1998). Compensatory Neutral Mutations and the Evolution of RNA. Genetica 102/103 91-101. Special issue published in book form as Mutation and Evolution. Eds. Woodruff, R.C., Thompson, J.N. Kluwer Academic Publishers, Dordrecht, Netherlands.
5. Higgs, P.G. (1996). Overlaps Between RNA Secondary Structures. Phys. Rev. Lett. 76, 704-707.
6. Morgan, S.R. & Higgs, P.G. (1996). Evidence for Kinetic Effects in the Folding of Large RNA Molecules. J. Chem. Phys. 105, 7152-7157.
7. Higgs, P.G. (1995). Thermodynamic properties of transfer RNA: a computational study. J. Chem. Soc. Faraday Transactions 91, 2531-40.
8. P.G.Higgs. RNA Secondary Structure: A Comparison of Real and Random Sequences. J. Phys. I 3, 43, (1993).
RNA secondary structure - the basics
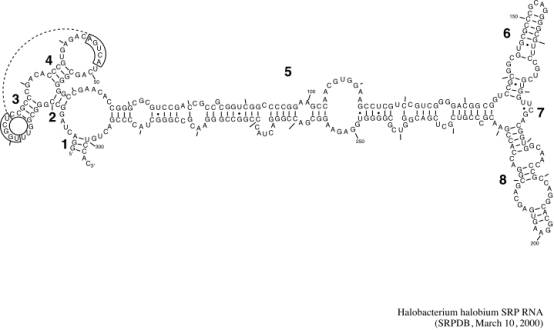
This is an example of a secondary structure of a medium-sized RNA molecule. It illustrates the typical pattern of base-paired regions (these are helices in 3 dimensions) separated by single stranded regions. We have borrowed this picture from the Signal Recognition Particle database. This is just one type of RNA molecule with complex secondary structure. Another web site with huge numbers of links to RNA software and information resources is the RNA World page.
RNA secondary structure is interesting to biologists because the structure is essential for the correct functioning of the molecule. A variety of bioinformatics algorithms are available to predict structures of given sequences using either thermodynamics/free energy minimization or by looking for conserved structures in sets of related sequences. RNA structure is of interest to physicists because it is complex problem in statistical mechanics with a rugged energy energy landscape (more details below).
I have recently completed a large-scale review of the methods used for RNA structure prediction and of statistical mechanics problems related to RNA [2].
Evolution of RNA via Compensatory Substitutions
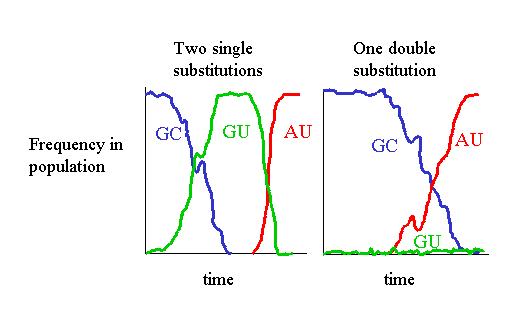
The secondary structure of RNA molecules such as ribosomal RNAs and transfer RNAs are strongly conserved during evolution yet the sequences vary considerably. Mutations accumulate in these sequences in such a way that the secondary structure is not disrupted - for example a GC pair in a helical region of a molecule in one species might be replaced by an AU pair in a related species. This involves a pair of substitutions occurring at two separate sites in the gene for the RNA molecule. This may either occur by two separate single substitutions, or by the compensatory substitution mechanism, in which the consensus sequence changes by two substitutions simultaneously. For a population genetics theory of this process, see [4].
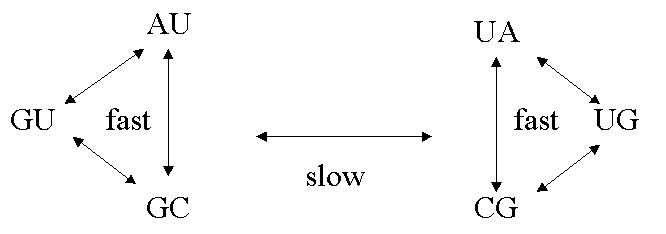
We recently completed a survey of paired regions of RNA in databases of tRNA, small sub-unit rRNA, and ribonuclease P RNA [2]. We showed that there is selection acting to stabilise the secondary structure, and hence that high-stability base-pairs are more frequent than low-stability ones. This pattern occurs in several of types of molecules, and across a range of organisms with differing content of GC and AU bases. From the sequence alignment data we are able to estimate relative rates of different types of substitution events: double transitions such as GC to AU, double transversions such as GC to CG, and single transitions like GC to GU. Thermodynamic stability of the base pairs also plays an important role in determining these rates. There are two groups of states in which rapid interchange occurs, whilst interchange between the two groups is slow [1,2].
Ribosomal RNA is one of the most important genes used in molecular phylogenetics. Almost all programs assume that different sites in a gene evolve independently. This is clearly not true in the paired regions of RNA (roughly half the sites in the sequence), hence there may be substantial bias in results. We have developed the PHASE package, which is a set of phylogenetic programs intended to account explicitly for RNA secondary structure. See the Phylogenetic Methods page for more details.
RNA Folding and Statistical Physics
We have been interested in the kinetics of the RNA folding process for some time. We have developed Monte Carlo programs that simulate the RNA folding process by formation and break-up of helices [2]. By comparing minimum free energy structures with biological structures obtained by comparative sequence analysis, we showed that there is evidence for kinetic effects in the folding of many large RNA sequences [6]. Large structures are composed of combinations of stable domains of medium size but are not necessarily minimum free energy structures. Evolution can act on RNA sequences, which means that the thermodynamic properties of real sequences are not the same as those of randomly generated sequences. In the case of tRNAs we have shown explicitly that the biological sequences are selected for stability of secondary structure [7,8] and that there are fewer alternative structures with energies close to the groundstate structure in real molecules than in random sequences.
RNA is also an example of what is known as a 'disordered system' in physics. This means that it has many alternative low energy structures that are separated from each other by high-energy barriers. We have worked recently on determining the size of these barriers, and the distribution of low-energy states in configuration space, in order to compare RNA with theoretical treatments of other models in statistical physics [3,5]. The alternative groundstates of a single random RNA can be clustered hierarchically according to the heights of the barriers between them. In the diagram below, dark squares are groups of similar structures separated by low barriers.
